Regulation of sumoylation by E2 enzyme (Ubc9) sumoylation
Lab Pichler
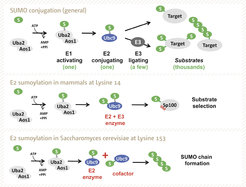
Figure 1 SUMO conjugation is regulated via E1, E2 and E3 enzymes (upper panel). E2 (Ubc9) sumoylation contributes to regulation of SUMO conjugation: in mammals Ubc9 is modified at Lysine 14 and this contributes to substrate selection (middle panel), in yeast Ubc9 is modified at Lysine 153, which inactivates the enzyme and turns it into a cofactor for SUMO chain assembly (lower panel).
Sumoylation is primarily regulated via E3 ligases and SUMO specific proteases because these enzymes mainly ensure substrate specificity. We characterized an alternative mechanism of regulation on the E2 level: E2 enzyme (Ubc9) sumoylation. This modification is conserved from yeast to mammals but involves structurally different sites of modification, suggesting distinct enzymatic consequences.
In mammalian cells, we found that N-terminal Ubc9 sumoylation enhances the affinity and modification of selected substrates in dependence of a non-covalent SUMO interaction motif (SIM) (Knipscheer, Klug et al., Mol Cell 2008, Figure 2, middle panel).
By contrast, C-terminal Ubc9 sumoylation in Saccharomyces cerevisiae results in E2 inactivation but turns this inactive Ubc9*SUMO into a cofactor for the unmodified Ubc9. Together, these enzymes cooperate in SUMO chain assembly, which is important for successful synaptonemal complex (SC) formation in yeast meiosis (Klug et al, Mol Cell 2013, lower panel).
We identified novel substrates for the mammalian sumoylated Ubc9 and are currently investigating how sumoylation regulates their biological functions in dependence of sumoylated Ubc9 and SUMO E3 ligases.
