Regulatory Circuitries of B Lymphopoiesis

B lymphopoiesis depends on the integration of extra-cellular signals by transcription factors that specify hematopoietic progenitors and allow for differentiation into highly-specialized effector cells. We are interested in understanding the molecular basis of B cell differentiation by dissecting the regulatory circuitries in which cell-type-specific transcription factors operate.
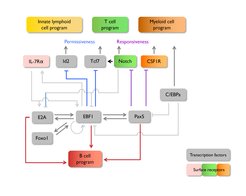
Toward this goal, we are studying the function of Early B cell Factor-1, EBF1, which is expressed in the early stages of the B cell lineage and in a subset of stromal cells in the bone marrow. Loss- and gain-of-function experiments indicated that EBF1 functions in a complex regulatory network with other transcription factors, in which positive feedback loops and cross-antagonism stabilize the establishment and maintenance of the B cell program. EBF1 is involved in activating genes associated with the B cell lineage and represses genes of alternative lineages.
In addition, EBF1 poises genes for expression at later stages of differentiation. Genome-wide and kinetic analysis of EBF1 occupancy indicated that EBFI binds naïve progenitor chromatin prior to the generation of accessibility and activation of gene expression. Currently, we are studying the molecular mechanisms by which EBF can fulfill its diverse roles as a lineage-determining factor.

