Regulation of neuron-specific RNA sequences
Hilgers Lab
Although a typical Drosophila 3’ UTR measures a few hundred base pairs, neuron-specific 3’ UTRs can be up to 17kb long. This extreme addition of sequence creates substantial potential for post-transcriptional regulation. We think that this extra layer of regulation is necessary for the mRNA to achieve a neuron-specific function.
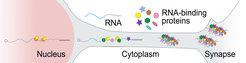
The role of RNA-binding proteins and RNA granules in neurological pathologies, in particular neurodegeneration, has been well described. Ultra-long 3’ UTRs, through their unusually high number of target sites for RNA-binding proteins, could represent a platform for regulated mRNA transport and translation, and may build a scaffold for mRNA granule assembly.
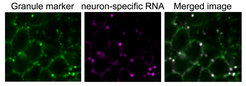
Applying imaging, neuron biochemistry, whole-genome analyses and functional genetics, we study how neuronal RNAs, in particular ultra-long 3’UTRs, are specifically regulated by RNA-binding proteins.

